
By showing examples with distinct CETSA behaviors, we aim to provide the scientific community with an overview of different scenarios to expect during CETSA experiments, especially for challenging, membrane bound targets. Membrane proteins are classified according to two different schemes. In single-pass transmembrane proteins, the polypeptide crosses only once (see example 1 in Figure 10-17 ), whereas in multipass transmembrane proteins, the polypeptide chain crosses multiple times (see example 2 in Figure 10-17 ). 1) How does a single-pass differ from a multi-pass transmembrane protein 2) What similarities/differences can you describe between an alpha helix vs. 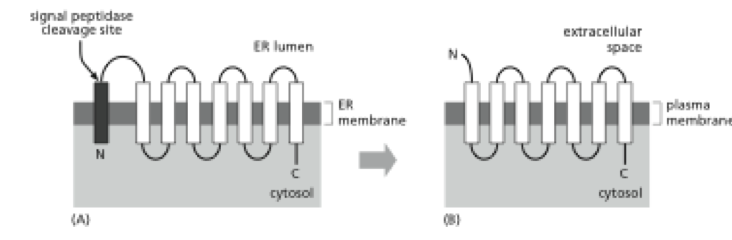
Our data demonstrated that using modified protocols with detergent extraction after the heating step, CETSA can reliably be applied to several membrane proteins of different complexity. Here, we present the application of live-cell CETSA to different classes of integral multipass transmembrane proteins using three case studies, the first showing a large and robust stabilization of the outer mitochondrial five-pass transmembrane protein TSPO, the second being a modest stabilization of SERCA2, and the last describing an atypical compound-driven stabilization of the GPCR PAR2. Compare and contrast the mechanism Ior localization ot single-pass vs multi- Pass transmembrane proteins In the ER Describe the processes of vesicle fuslon. Both are single membranespanning synthesized peptides with fewer than 36 residues. The cellular thermal shift assay (CETSA) has been introduced as a powerful label-free method to assess target engagement in physiological environments. The computational design of transmembrane proteins with more than one. Demonstration of target binding is a key requirement for understanding the mode of action of new therapeutics.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
